- Home
- News
- Spotlight on Science
- Cryo-EM structures...
Cryo-EM structures of the chikungunya virus nsP1 reveal capping rings as functional gates of membranous viral factories
12-02-2021
Cryo-electron microscopy structures of the chikungunya virus nsP1 protein, obtained at beamline CM01, reveal the structural basis of the coupling between membrane binding, oligomerisation and allosteric activation of the capping enzyme, which underpins the association of the viral replication machinery with virus-induced membranous organelles within host cells.
Studies of the replication complexes of positive-stranded RNA viruses have been hampered by their membrane association, making them elusive for structural studies. Researchers from Marseille (Architecture et Fonction des Macromolecules Biologiques laboratory, CNRS-AMU) managed to isolate a membrane-bound complex from one of these replication complexes; the chikungunya virus RNA capping protein, nsP1.
For decades, nsP1 was studied in its monomeric form, as it comes from E. coli recombinant expression. Using eukaryotic expression systems and detergents, homogeneous and very active ring-shaped complexes were obtained, in contrast with the poorly active monomeric nsP1. The complexes were prepared for single particle cryo-EM structure determination with the help of a researcher from Madrid, before being brought to the ESRF and imaged using the Titan Krios electron microscope at beamline CM01, delivering extraordinary data that allowed the 3D reconstruction of the nsP1 rings at 2.6-Å resolution.
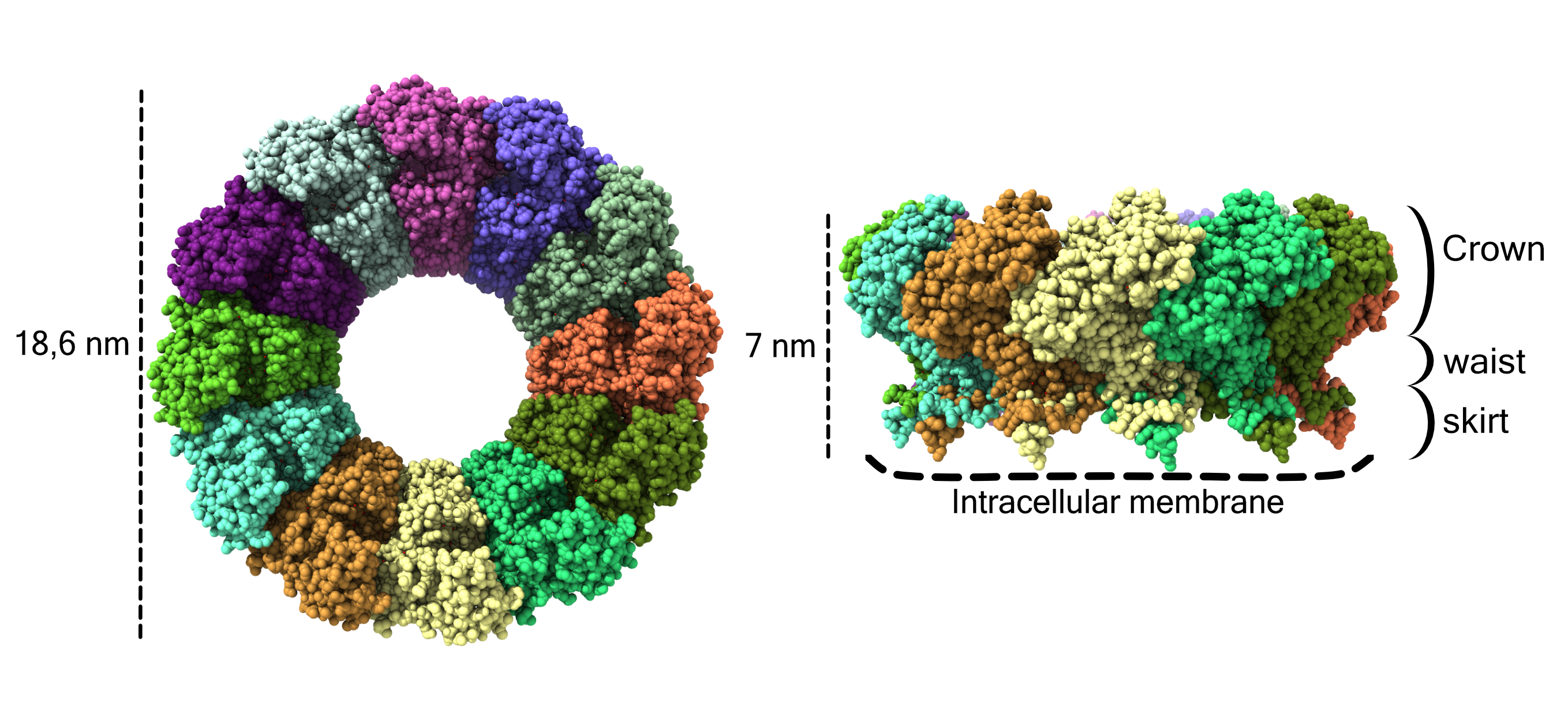
Fig. 1: Atomic structure of the pores of the chikungunya nsP1 virus with the different monomers shown in different colours. Each atom is represented as a sphere. On the left side of the image the pore is shown from the cytoplasmic view and on the right from a side view. The dimensions of the pore are indicated on the left, the crown, waist and skirt regions are indicated on the right and the contact sites with the membrane are indicated at the bottom by dashed lines.
The resulting cryo-EM structure revealed that nsP1 forms homo-dodecameric rings, defining three important regions in its architecture (Figure 1). The crown of the ring harbours the capping domains, which stack against each other to stabilise their active sites conformation. The capping domains have similar folding to homologous SAM-dependent methyl transferases, enzymes broadly conserved in all kingdoms of life, and that are involved in RNA labelling. The peculiarity of this enzyme is that it is able both to methylate and transfer the CAP structure to the RNA, essential for its stability and recognition by the ribosomes.
Below the crown, the complex narrows to the waist, the region defining an internal pore from where the viral RNA could access the interior of viral replication organelles and from where the newly synthetised viral RNAs would be expelled to the cytoplasm properly labelled by the capping enzyme. Finally, a twelvefold spiked skirt underneath the waist is mediating a very strong interaction with membranes in a monotopic fashion (without traversing the entire lipid bilayer), but reaching a depth of more than 10 Å into the lipid bilayer. The cone-shape of the spikes and the surrounding charge distribution are indicative of a very strong potential for membrane deformation.
Thus, in its singular architecture, the complex combines the ability to bind and bend membranes, generate a membrane pore and trigger the enzymatic activity and form a molecular platform for the attachment of other viral factors such as the polymerase [1] – and all this with only a small (61 KDa) nsP1 protein.
The structure of alphavirus capping rings provides the first structural information, at high resolution, of the ways RNA viruses replicate and transcribe their genomes in their membranous environment. For example, all alphaviruses (e.g., Semliky forest virus or Venezuelan equine encephalitis virus), Rubella virus, hepatitis E virus or other related flaviviruses such as Dengue or hepatitis C, or even the threatening coronavirus, carry out replication within membrane organelles. Still, the available information about their replication machineries neglects their membrane association, thus current understanding of the way they work limits capacity to, for instance, develop effective antivirals.
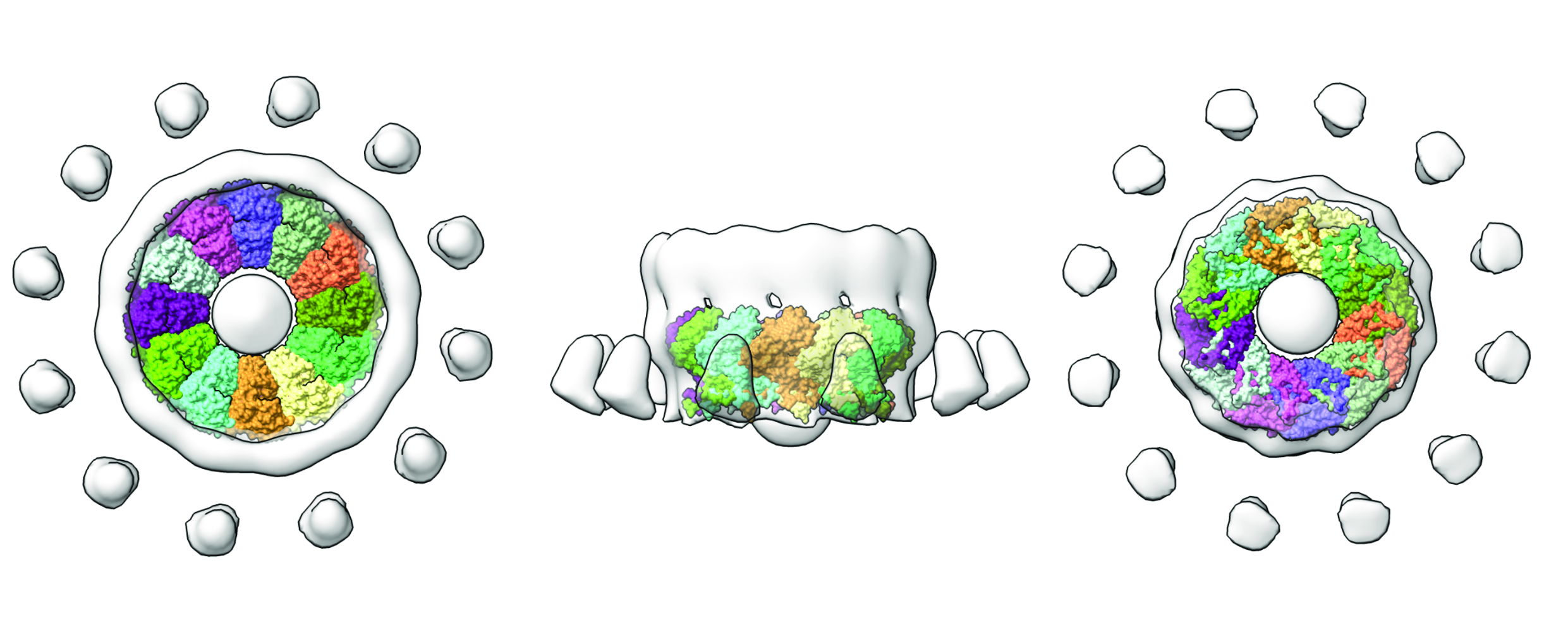
Fig. 2: nsP1 complex (coloured by protomer) superposed into the tomography map reconstructed from FHV growing spherules (light grey surface). On the left, the complex is shown from the cytosolic view, in the middle from the lateral view, and on the right from the interior of the viral organelle. The figure was made with ChimeraX.
With this structure in hand, it is now possible to better interpret the existing structures of replication complexes at much lower resolution obtained by cryo-electron tomography. This is the case for the structure of the Flock House Virus crown-like structures observed at the necks of spherule-shaped viral organelles [2]. Superposition of this tomographic reconstruction with the nsP1 capping rings illuminates the conservation of these structures between a broad group of positive-stranded RNA viruses (Figure 2). Other tomographic reconstructions of viral factories in coronavirus-infected cells, despite notable differences in the architecture, also show the oligomerisation of replication complexes into macromolecular pores [3].
Thus, the structure of nsP1 capping rings opens a new path for the understanding of viral replication within membrane organelles. This will uncover essential functional aspects of RNA virus biology, possibly leading to the creation of more effective antiviral drugs to contain and fight against future pandemics of emerging infectious diseases by RNA viruses.
Principal publication and authors
Capping pores of alphavirus nsP1 gate membranous viral replication factories, R. Jones (a), G. Bragagnolo (a), R. Arranz (b), J. Reguera (a,c), Nature 589, 615-619 (2021); https://doi.org/10.1038/s41586-020-3036-8.
(a) CNRS, AFMB UMR 7257, Aix-Marseille Université, Marseille (France)
(b) National Center of Biotechnology, CSIC, Madrid (Spain)
(c) INSERM, AFMB UMR7257, Marseille (France)
References
[1] M.K. Pietila, K. Hellstrom, T. Ahola, Virus Research 234, 44-57 (2017).
[2] N. Unchwaniwala et al., PNAS 117, 18680-18691 (2020).
[3] G. Wolff et al., Science 369, 1395-1398 (2020).



